Database Statistics
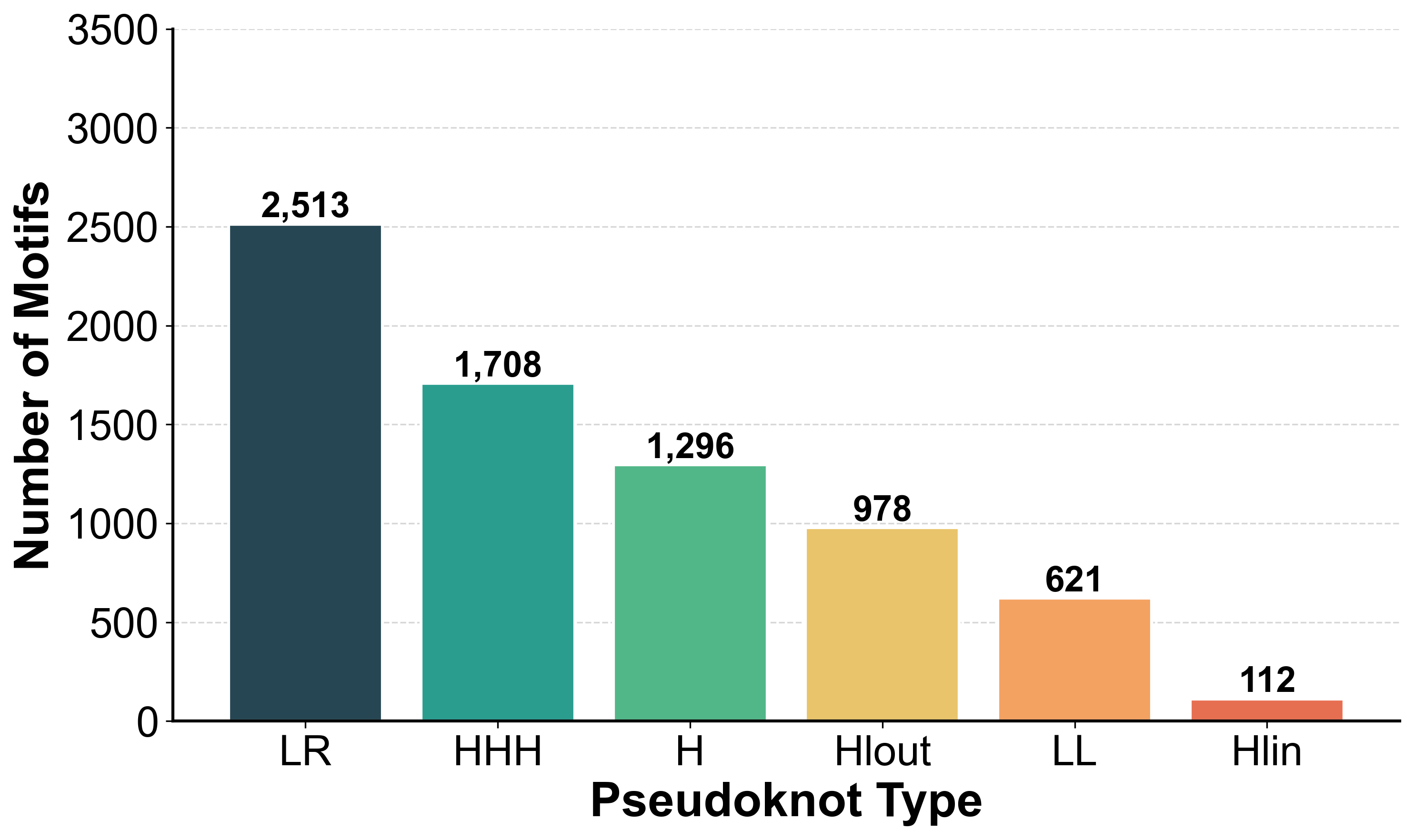
Distribution of motif types and sizes in ATLAS
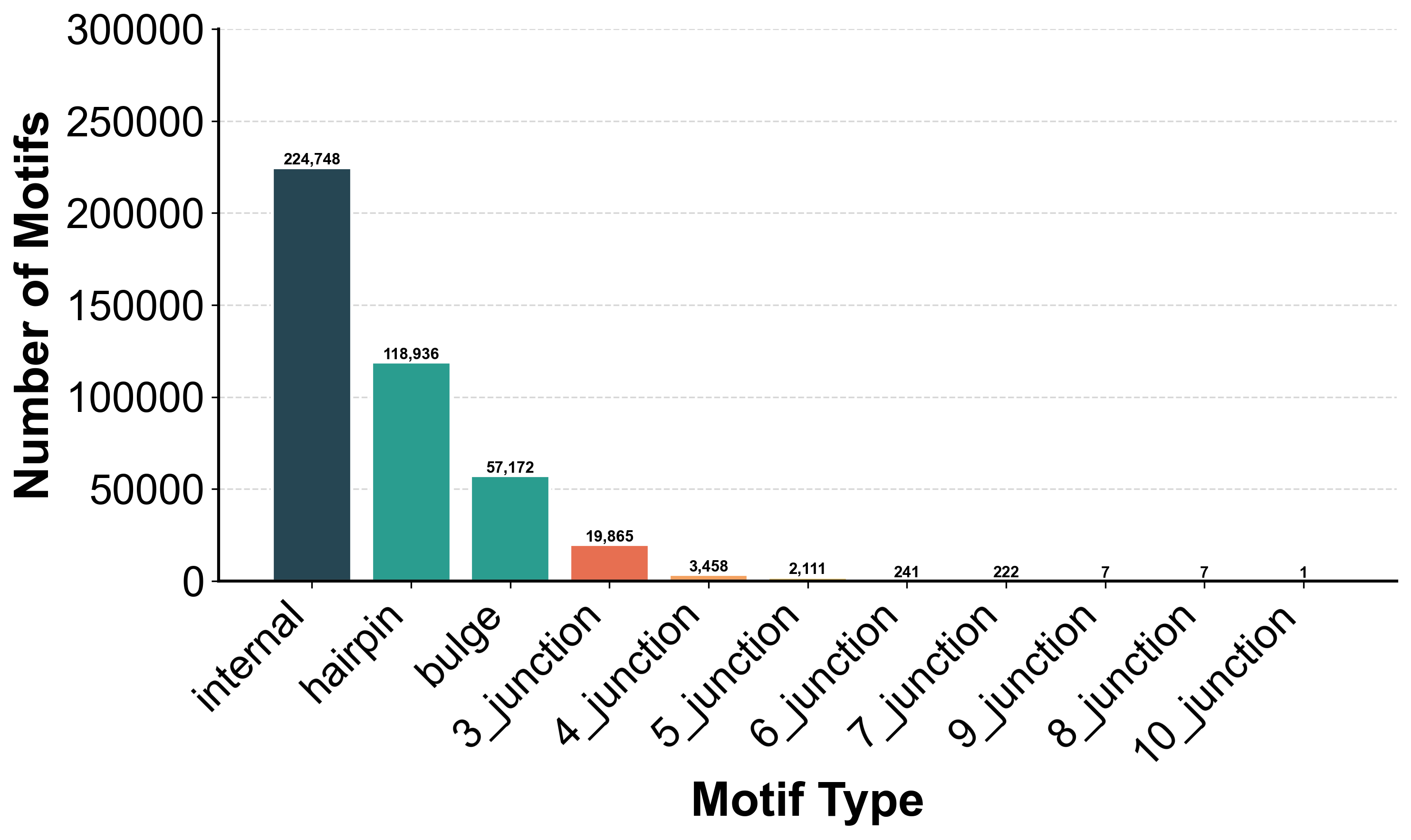
Motif Types
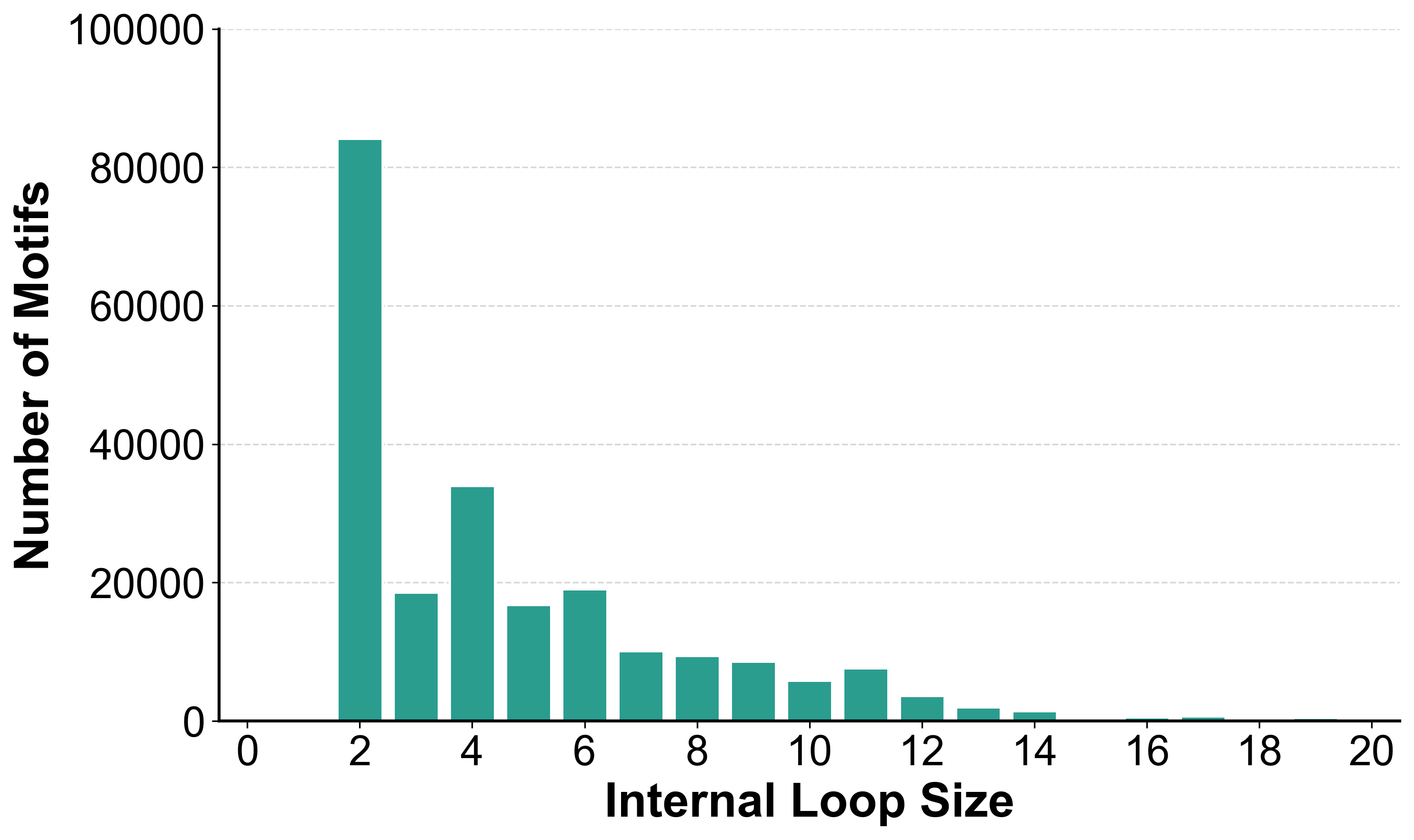
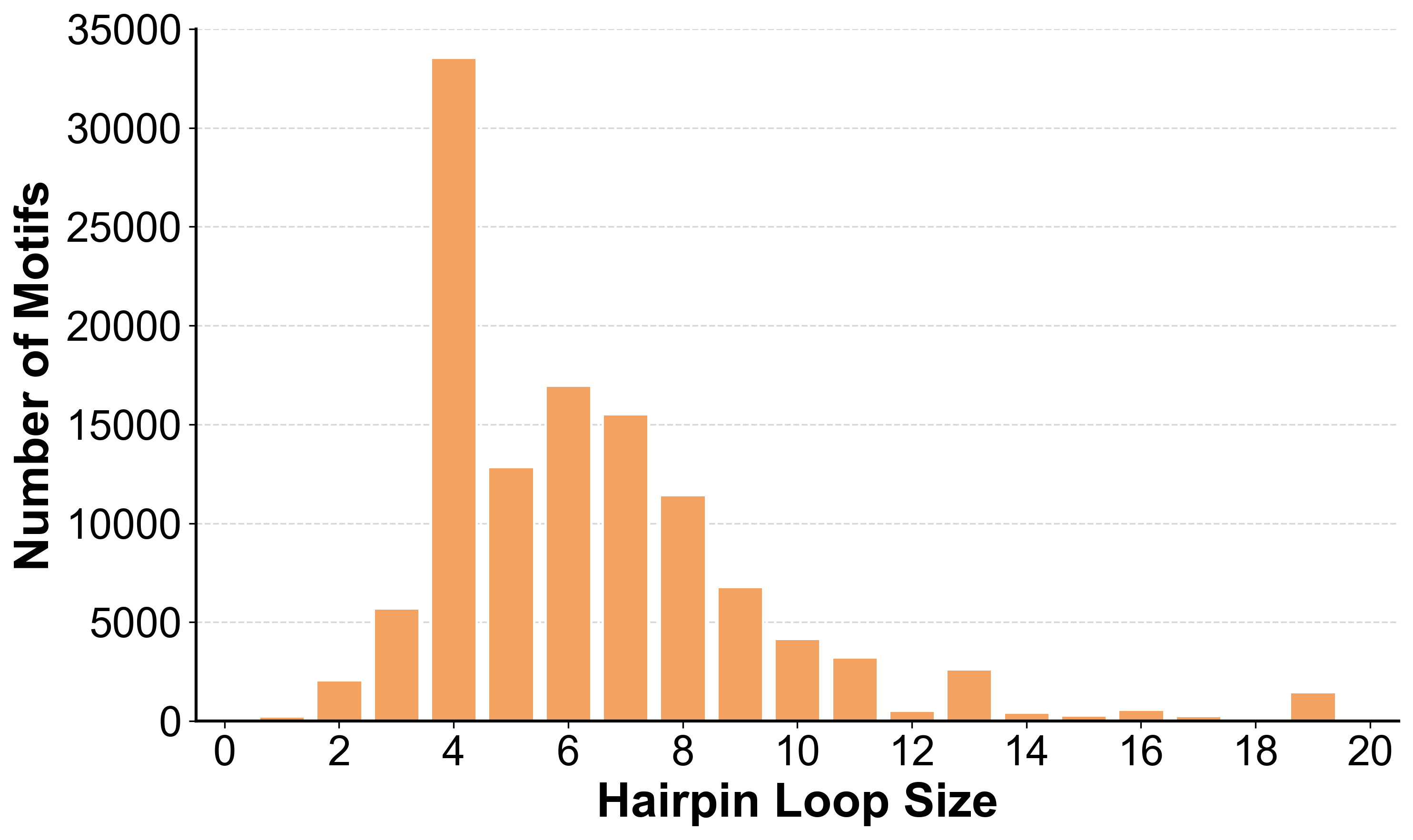
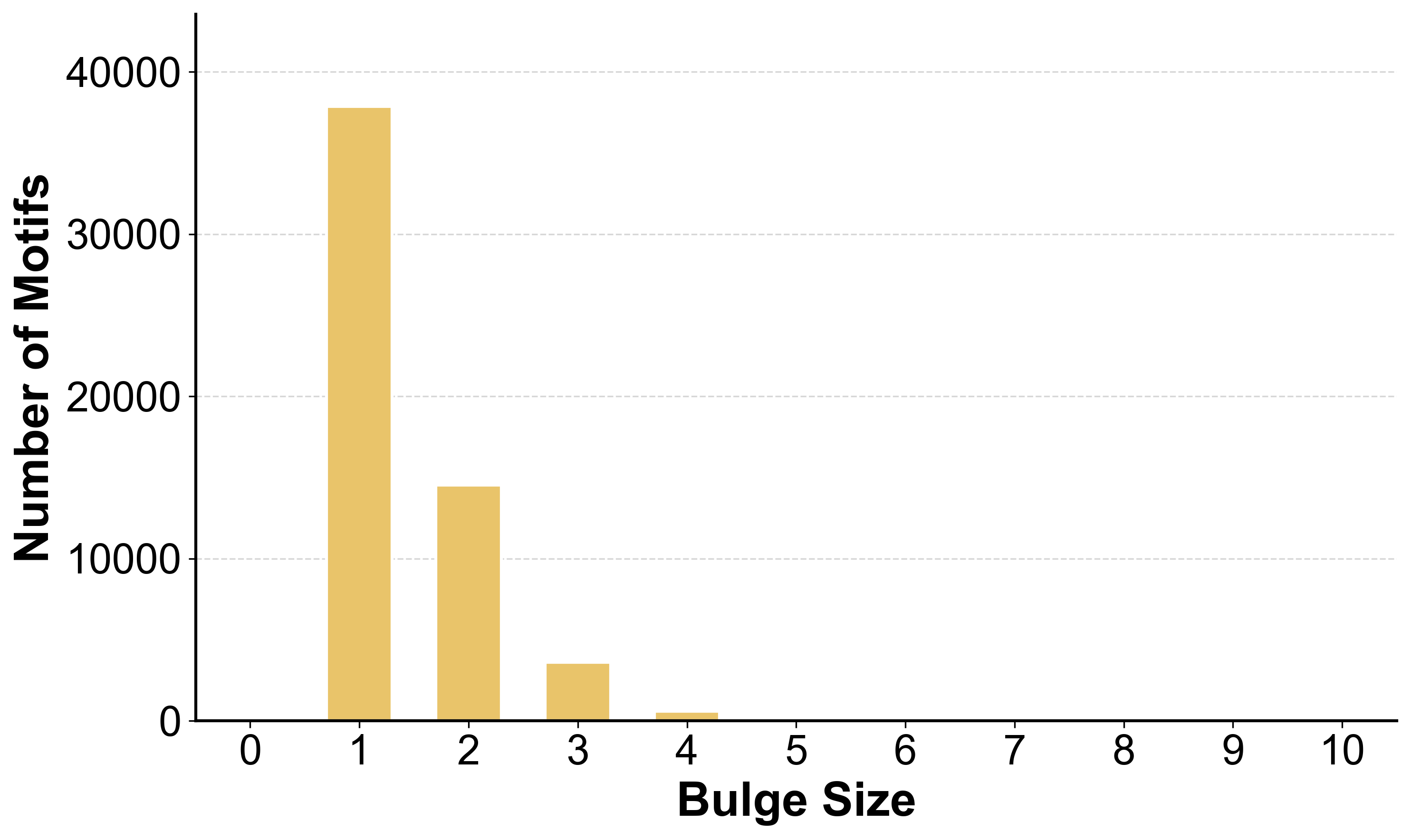
Size Distribution
Advanced Template Library for Assembly and Structure
ATLAS is a comprehensive 3D RNA motif library that enables researchers to search for RNA structural motifs in the Protein Data Bank. Using graph-based algorithms, ATLAS extracts and catalogs over 433,000 RNA motif structures including hairpins, internal loops, bulges, multiway junctions, and pseudoknots. The database provides atomic-level 3D coordinates for accurate structural analysis and RNA research.
We have designed a user-friendly website dedicated to RNA motif search, utilizing a graph-based search algorithm to extract 3D RNA motif structures.
ATLAS offers two powerful search methods for RNA structural motifs from the Protein Data Bank.
Over 433,000 motifs from PDB: 52% Internal loops, 27% Hairpins, 13% Bulges, 6% Junctions, 2% Pseudoknots
Search pre-indexed RNA motifs from ATLAS database by selecting motif type (hairpin, internal loop, bulge, multiway junction, or pseudoknot) and motif size (number of unpaired nucleotides). Results include 3D atomic coordinates and graph representations from the Protein Data Bank (PDB).
Draw your own RNA motif structure using an interactive graph-based interface. The search uses subgraph isomorphism algorithm to find matching patterns in RNA structures. Supports non-Watson-Crick interactions for comprehensive structural analysis.
Distribution of motif types and sizes in ATLAS





For questions, support, or feedback about the ATLAS database, please contact:
Nikolay V. Dokholyan, PhD
Professor
Department of Neurology
Department of Neuroscience
Department of Biomedical Engineering
Department of Microbiology, Immunology, & Cancer Biology
University of Virginia